Christine Spang blogs here about free software hacking and her adventures and endeavors.
Have a look at the most recent posts below, or browse the tag cloud on the right. An archive of all posts is also available.
Non-blog things of Christine's can be found on her homepage.
Last week, in the spirit of always getting better, I asked Twitter how to practice saying "um" less while speaking. Here's what I learned.
Most people suggested to practice intentionally inserting pauses instead of saying "um". Various strategies include voice acting classes, having a friend aid with a buzzer (presumably to help you notice), and to record practice sessions as well as the real deal.
Nagle suggested that meditation helped him, which makes sense to me as I've experienced meditation helping me be more mindful in many different areas of my life, including mundane things like noticing that I've touched something that's probably not that clean and remembering not to touch my face until I wash them. Wilfully directing mindfulness toward noticing how I speak is something I'm really stoked about, though it's easy to get swept away in a default "performance mode" in front of a crowd.
Hendrick pointed out that Cognitive Behavioural Therapy is the principle behind changing speech behaviour and is a great starting point for more research.
This is going to be really useful next time I'm practicing for a talk!
Thanks @zmagg,
@hendricklee,
@bcrypt,
@ebroder,
@mscain,
and @nagle5000 for playing. 
PyCon 2014 happened. (Sprints are still happening.)
This was my 3rd PyCon, but my first year as a serious contributor to the event, which led to an incredibly different feel. I also came as a person running a company building a complex system in Python, and I loved having the overarching mission of what I'm building driving my approach to what I chose to do. PyCon is one of the few conferences I go to where the feeling of acceptance and at-homeness mitigates the introvert overwhelm at nonstop social interaction. It's truly a special event and community.
Here are some highlights:
- I gave a tutorial about
search,
which was recorded in its entirety... if you watch this video, I
highly recommend skipping the hands-on parts where I'm just walking
around helping people out.

- I gave a talk! It's called Subprocess to FFI, and you can find the video here. Through three full iterations of dry runs with feedback, I had a ton of fun preparing this talk. I'd like to give more like it in the future as I continue to level up my speaking skills.
- Allen Downey came to my talk and found me later to say hi. Omg amazing, made my day.
- Aux Vivres and Dieu du Ciel, amazing eats and drink with great new and old friends. Special shout out to old Debian friends Micah Anderson, Matt Zimmerman, and Antoine Beaupré for a good time at Dieu du Ciel.
- The Geek Feminism open space was a great place to chill out and always find other women to hang with, much thanks to Liz Henry for organizing it.
- Talking to the community from the Inbox booth on Startup Row in the Expo hall on Friday. Special thanks for Don Sheu and Yannick Gingras for making this happen, it was awesome!
- The PyLadies lunch. Wow, was that amazing. Not only did I get to meet Julia Evans (who also liked meeting me!), but there was an amazing lineup of amazing women telling everyone about what they're doing. This and Noami Ceder's touching talk about openly transitioning while being a member of the Python community really show how the community walks the walk when it comes to diversity and is always improving.
- Catching up with old friends like Biella Coleman, Selena Deckelmann,
Deb Nicholson, Paul Tagliamonte, Jessica McKellar, Adam Fletcher, and
even friends from the bay area who I don't see often. It was hard to
walk places without getting too distracted running into people I knew,
I got really good at waving and continuing on my way.

I didn't get to go to a lot of talks in person this year since my personal schedule was so full, but the PyCon video team is amazing as usual, so I'm looking forward to checking out the archive. It really is a gift to get the videos up while energy from the conference is still so high and people want to check out things they missed and share the talks they loved.
Thanks to everyone, hugs, peace out, et cetera!
Today I gave a tutorial at PyCon 2014 entitled Search 101: An Introduction to Information Retrieval.
It was an experiment of sorts: the first workshop I've run primarily by myself, my first tutorial at PyCon, my first paid teaching gig. It was an opportunity to take some of the lessons I learned from teaching the Boston Python Workshop and apply them to a new situation.
The material itself is a distillation of many hours of frustration with the documentation for various open source search engine libraries, frustration that they didn't tell me where to start or about the big picture, they just jumped straight into the details.
Here's what worked:
- IPython Notebook. Oh em gee. I started writing the class's handout using IPython Notebook because it was a simple way to easily embed syntax-highlighted code into a markdown document that was viewable in a browser. Not only was it a super quick and fun way to write the handout, but many students used the interactive execution features to play around with the example code.
- Not having a paper handout. Saved trees, printing hassle, and no one seemed to mind.
- Putting everything in a git repo... git is sufficiently ubiquitous these days that students didn't really have trouble getting a copy, and appreciated having everything in one place, with simple setup instructions. I brought a clone of the repo on a USB stick as a backup plan.
Here's what caused problems:
- Mostly, the IPython dependency
pyzmq, which requires compilation. I don't know what the current landscape is for Python distribution, but installing these libraries throughpipis still a pain. I've heard rumour that more ubiquitous wheels may solve this in the future. - Some people aren't used to using
virtualenveverywhere. Even seeing that, I still think it's worth the confusion to put it forth as the recommended setup method.
Intermediate students are a different crowd than beginners. There was less of an air of discovery in the room, though I organized the class around open-ended tasks. Since the material allowed for folks to take it in the direction of their interest, I found it a bit difficult to gauge whether people were following or not. Overall though, everyone was attentive and studious. I had fun.
Ruben and Stuart, the PyCon tutorial organizers, had logistics running
super smoothly, AV, lunch, everything. Thanks for that you guys, you
rock.  And thanks as well to my helpers: Leo, the tutorial host, Eben,
my TA, and Roberto, on AV. It's impossible to pay adequate attention to
20+ people as a single person, couldn't have done a decent job without
y'all.
And thanks as well to my helpers: Leo, the tutorial host, Eben,
my TA, and Roberto, on AV. It's impossible to pay adequate attention to
20+ people as a single person, couldn't have done a decent job without
y'all. 
At Inbox we're using Phabricator to review all code going to production. It's a great tool that enables us to easily learn from each other and increase the quality of the code we're writing.
Phabricator is pretty easy to install, but it requires running a bunch of
daemons that perform background tasks. It provides a wrapper script called
phd which, when invoked, spins up the default set of daemons and backgrounds
them. Unfortunately, since it backgrounds the processes, it doesn't play well
with the excellent Supervisor, which allows one to
automatically bring the daemons up after a reboot or if they crash, since
supervisor requires managed processes to stay in the
foreground.
If you try to use the obvious phd launch under supervisor... wat wat.
$ supervisorctl status
phd FATAL Exited too quickly (process log may have details)
The process error log ends up looking like:
phd start: Unable to start daemons because daemons are already running.
You can view running daemons with 'phd status'.
You can stop running daemons with 'phd stop'.
You can use 'phd restart' to stop all daemons before starting new daemons.
supervisor starts the daemons, but it can't tell that they're running
because they automatically background themselves!
There's a debug mode to phd, but running in production we don't necessarily
want /var/log to fill up with mountains of debug spew.
Instead, stick the following in your supervisor configuration (on
Debian/Ubuntu, paste into the new file /etc/supervisor/conf.d/phd.conf):
[program:PhabricatorRepositoryPullLocalDaemon]
command=/srv/phabricator/scripts/daemon/phd-daemon PhabricatorRepositoryPullLocalDaemon --phd=/var/tmp/phd/pid
autostart=true
autorestart=true
stdout_logfile=/var/log/supervisor/PhabricatorRepositoryPullLocalDaemon.log
stderr_logfile=/var/log/supervisor/PhabricatorRepositoryPullLocalDaemon_err.log
[program:PhabricatorGarbageCollectorDaemon]
command=/srv/phabricator/scripts/daemon/phd-daemon PhabricatorGarbageCollectorDaemon --phd=/var/tmp/phd/pid
autostart=true
autorestart=true
stdout_logfile=/var/log/supervisor/PhabricatorGarbageCollectorDaemon.log
stderr_logfile=/var/log/supervisor/PhabricatorGarbageCollectorDaemon_err.log
[program:PhabricatorTaskmasterDaemon1]
command=/srv/phabricator/scripts/daemon/phd-daemon PhabricatorTaskmasterDaemon --phd=/var/tmp/phd/pid
autostart=true
autorestart=true
stdout_logfile=/var/log/supervisor/PhabricatorGarbageCollectorDaemon1.log
stderr_logfile=/var/log/supervisor/PhabricatorGarbageCollectorDaemon1_err.log
[program:PhabricatorTaskmasterDaemon2]
command=/srv/phabricator/scripts/daemon/phd-daemon PhabricatorTaskmasterDaemon --phd=/var/tmp/phd/pid
autostart=true
autorestart=true
stdout_logfile=/var/log/supervisor/PhabricatorGarbageCollectorDaemon2.log
stderr_logfile=/var/log/supervisor/PhabricatorGarbageCollectorDaemon2_err.log
[program:PhabricatorTaskmasterDaemon3]
command=/srv/phabricator/scripts/daemon/phd-daemon PhabricatorTaskmasterDaemon --phd=/var/tmp/phd/pid
autostart=true
autorestart=true
stdout_logfile=/var/log/supervisor/PhabricatorGarbageCollectorDaemon3.log
stderr_logfile=/var/log/supervisor/PhabricatorGarbageCollectorDaemon3_err.log
[program:PhabricatorTaskmasterDaemon4]
command=/srv/phabricator/scripts/daemon/phd-daemon PhabricatorTaskmasterDaemon --phd=/var/tmp/phd/pid
autostart=true
autorestart=true
stdout_logfile=/var/log/supervisor/PhabricatorGarbageCollectorDaemon4.log
stderr_logfile=/var/log/supervisor/PhabricatorGarbageCollectorDaemon4_err.log
Make sure you're not running phd manually, restart the supervisor service
(sudo service supervisor restart on Debian/Ubuntu) and you should be good to
go.
(Much thanks to Evan Priestley for the quick
support and explanation on #phabricator.) Note that this solution is
officially not supported and could break.
Installing fresh hot Debian 7.0 on a shiny new ThinkPad X1 Carbon laptop turns out to be easy as cake. You just need to make sure to grab the wifi firmware from unstable instead of the all-in-one firmware tarballs, which contain a version that is missing a couple required files.
wget http://cdimage.debian.org/debian-cd/7.0.0/multi-arch/iso-cd/debian-7.0.0-amd64-i386-netinst.iso
dd if=debian-7.0.0-amd64-i386-netinst.iso of=/dev/sdb
(Make sure /dev/sdb is really the usb stick you want to overwrite with the installer!)
wget http://ftp.us.debian.org/debian/pool/non-free/f/firmware-nonfree/firmware-iwlwifi_0.38_all.deb
And put that on a second usb stick for the installer to load the firmware off of.
As far as I can tell, everything works. (Did not mess around with the fingerprint reader, don't care.)
I just donated $500 to OpenHatch. Here's why you should donate too:
- Diversity in open source matters. We can't keep making the software the world runs on without involving people of all sorts, from all backgrounds.
- OpenHatch is run by community members who I've known for years and trust. They care about data-driven effectiveness and are always getting better at what they do.
- A rising tide floats all boats. More contributors == more awesome.
- If you donate before December 24th, your donation makes twice the difference.
Diversity and education initiatives are the reason I'm a part of the free and open source software community today. (Thanks, Debian Women.)
You don't have to donate $500 to make a difference. $5, $10, $25— from a hundred people—all adds up.
I'm studying technical leadership right now, out of necessity. My company is growing, and too many startups leave organizational matters in the chaos state until they're incredibly difficult to fix. If we can start designing the organization how we want it from the beginning, we can both make ourselves happier and more effective right now and greatly ease our troubles down the line.
My friend Matt Zimmerman recommended me a great book by Jerry Weinberg, Becoming a Technical Leader. Here's a gem from the book that resonated with me:
There are many technical workers who enjoy wandering so much that, like Alice in Wonderland, they don't much care where they go, so long as they get somewhere. Computer programmers call this process "hacking."
This is an apt description of many of the hackers I know, who are happy as long as the problem is technically interesting. But in order to accomplish anything, we need to place the act and experience of hacking within a system that directs what we're working on towards a goal.
More on this as the research and practical experience continue.
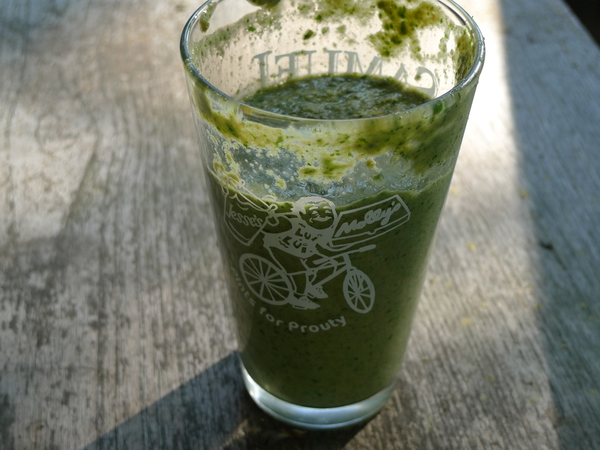
I made this smoothie this morning in a frantic attempt to use up some of my now-wilting CSA greens. The minute I tasted it I was blown away by how well it turned out!
Here's the recipe:
- 1 red apple
- 1 banana
- 1 bunch cilantro
- 3 leaves kale
- enough unsweetened soymilk to blend
Yes, I put in an entire bunch of cilantro. Result? A refreshing kick.
It's possible I could have stuffed in some more kale, or some spinach, or some salad greens, but after tasting it after the first blend, I was so happy I couldn't bring myself to experiment on it.
As a vegan and a female, I often wonder whether I'm getting proper nutrition—especially when it comes to things like iron, vitamin D, calcium, and B12, since these important nutrients are found in high concentrations in animal products. While I can make guesses as to what I'm not getting enough of based on how I feel, I've never really felt certain whether I really need this supplement, or whether I ought to be eating more lentils or what. Because the real way to figure that out is to get a blood test, and I'm not a doctor and getting blood tests for personal reasons in the US sounds expensive and like a lot of work.
 |
That's why InsideTracker caught my attention. It's a service where you sign up for a blood draw, and they perform a lab analysis on your blood for what they call "biomarkers"; basically a measure of the level of certain nutrients in your blood. Then their websites gives you nutrition suggestions to improve your results. As a person with a science background, and someone who wants to be in charge of my health and maximize my well-being and energy, it sounded perfect.
I gave it a shot. Here are my initial impressions.
pros
- Simple product choice. They only have two products, and are very honest about the fact that this is because the B12 and D analyses are a significant additional expense. I appreciated this explanation, and still opted for the "Fitness Plus" product because I'm particularly interested in those two markers.
- 24-hour turnaround from blood draw to data on the website. Wow, I'm impressed.
- God I love data. It's enough to get me excited about needles.
cons
- Their website still has a few rough edges. I got distracted in the middle of the signup process on the first go, and ended up in a multi-day email back and forth with someone on the team there who kept forwarding my problems getting my password reset to the engineering team. And I never got any notification that my problem was finally solved. A suggestion to the team: treat your early customers like gold, especially when they bring a problem to you rather than walking away. They'll be your biggest asset if you do.
- They sometimes have marketing deals, but they seem poorly put together. They were running a promotion for a while for free home blood draw, which they normally charge $80 for, but I either just barely missed it because I was dealing with account problems, or couldn't figure out how to actually apply for the promotion. So I ended up trekking from Cambridge to Newton to visit a LabCorp office instead.
results
My results were part unsurprising, part surprising. I have low blood iron ("ferritin" and "hemoglobin"), low vitamin D (thanks Boston winter), and elevated B12. I'm guessing the B12 is a combination of my nutritional yeast obsession, fortified cereals, and the multivitamin I take daily.
action plan
I came up with a few things I plan to do based on my results.
- phase in a vitamin D supplement, at least until it gets warm enough to get sunlight exposure
- buy a big bag of red lentils and use them as a base to add vegetables to for meals
- phase out the multivitamin I currently take daily, and cut back on breakfast cereal (which I don't usually eat for breakfast, but rather as a snack when I'm feeling too lazy or too hungry to cook or in the mood for something a bit desserty)
- increase my leafy greens intake
I'll aim for the lower end of the 3-6 months recommendation for my next blood test. I particularly want to make progress on the low blood iron levels.
I wonder how much blood levels of these "biomarkers" fluctuate on, say, a day-to-day basis. I would love to be able to get faster feedback, and I wonder how much the immediate context of when you get your blood drawn affects results (despite the required 12-hour fast). I was on my period when I got my blood drawn this time. Does that affect my results? Unfortunately, getting more frequent blood draws at this point is prohibitively expensive.
Even the 3-6-monthly blood draw is a fairly big expense, but if you're someone like me who can spring for a $30 monthly supplement that you might not actually need, the cost savings from the feedback might make it cheap. And if it makes you healthier, that may be worth the cost.
We'll see what I think after I get my next test in a few months.
In a little town called New Paltz, not far from a beautiful series of cliffs called the Shawangunks, exists a wonderful little vegan cafe called the Karma Road Cafe.
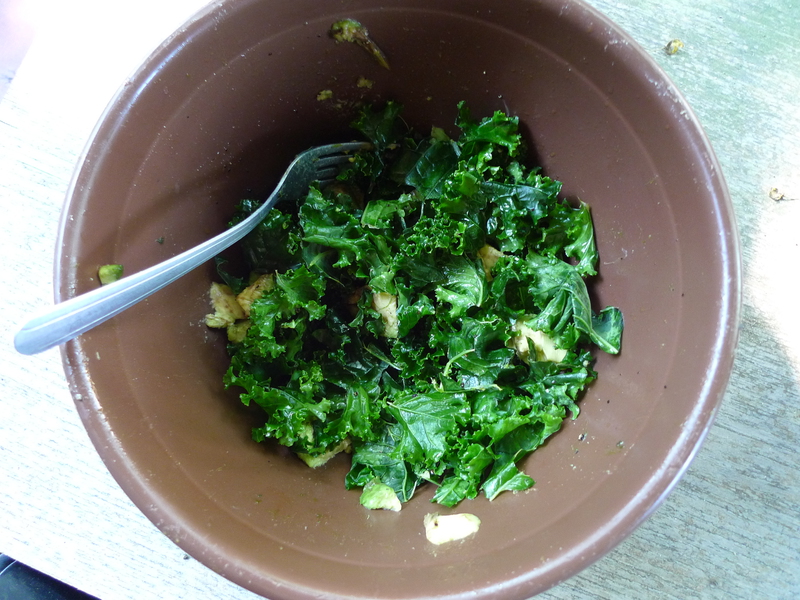
After a climbing trip one weekend, I reinvigorated my exhausted body with a wonderful thing there that they called the Kalecado salad. I recently attempted to reproduce it at home using fresh Red Fire Farm green curly kale from my weekly CSA share.
Here's what's in it:
- 1 bunch fresh green curly kale
- 1 ripe avocado
And for the dressing:
- equal parts olive oil and lemon juice (I used approx 3 tbsp each)
- salt and fresh-ground black pepper to taste
Chop or tear the kale into small pieces. Mix the dressing together and then mix it with the kale, using your hands to rub it in. Top with diced avocado.
I'm sure avocado was in a more pureed form in the original salad, and I don't recall the black pepper, but it came out delicious even if the details aren't quite right.
Fresh kale really is better raw.
This blog is powered by ikiwiki.

